Single-cell RNA sequencing workflow
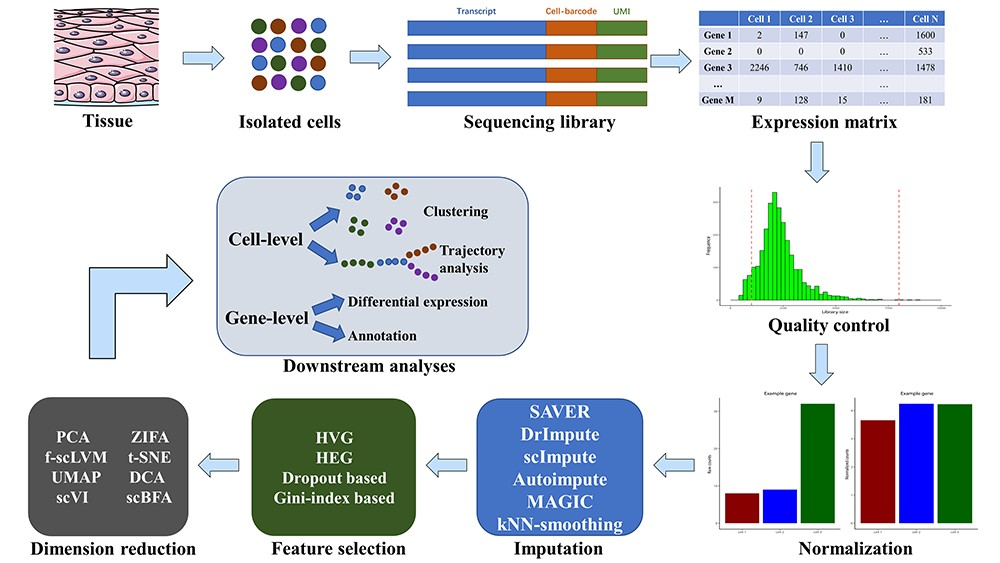
Single cells are first isolated from biological tissue, and mRNA is captured and amplified to obtain the sequencing library. After sequencing, the gene expression matrix is processed by QC, normalization, imputation, feature selection and dimension reduction in a step-by-step process. Finally, the processed expression matrix can be used for cell-level and gene-level downstream analyses.
This brief workflow was quoted from:
https://academic.oup.com/bib/article/22/4/bbaa314/6034054(Zhang et al., 2020)
If you would like to learn more about the process and details of bioinformatics analysis for scRNA-seq, you can read the following review papers, which we sincerely hope will help you in your work:
https://academic.oup.com/bib/article/22/4/bbaa314/6034054(Zhang et al., 2020)
https://www.nature.com/articles/s12276-018-0071-8(Hwang et al., 2018)
And two specially for plant science:
https://linkinghub.elsevier.com/retrieve/pii/S1360-1385(21)00247-8(Denyer & Timmermans, 2022)
https://linkinghub.elsevier.com/retrieve/pii/S1674205220303798(Shaw et al., 2021)
References
Denyer, T., & Timmermans, M. C. P. (2022, Jan). Crafting a blueprint for single-cell RNA sequencing. Trends Plant Sci, 27(1), 92-103. https://doi.org/10.1016/j.tplants.2021.08.016
Hwang, B., Lee, J. H., & Bang, D. (2018, Aug 7). Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med, 50(8), 1-14. https://doi.org/10.1038/s12276-018-0071-8
Shaw, R., Tian, X., & Xu, J. (2021, Jan 4). Single-Cell Transcriptome Analysis in Plants: Advances and Challenges. Mol Plant, 14(1), 115-126. https://doi.org/10.1016/j.molp.2020.10.012
Zhang, Z., Cui, F., Wang, C., Zhao, L., & Zou, Q. (2020). Goals and approaches for each processing step for single-cell RNA sequencing data. Briefings in Bioinformatics, 22(4). https://doi.org/10.1093/bib/bbaa314