1. Introduction: PscOA
PscOA (a database of Plant marker genes derived from Original Analyses of Single-Cell transcriptomic studies)
Single-cell transcriptomics offers a high-resolution analysis of the transcriptome at the single-cell level, uniquely positioning it to study cellular heterogeneity that bulk transcriptomics cannot achieve. Nevertheless, the advancement of plant science in this field lags behind that of animal science, due in large part to the scarcity of suitable marker genes.
To perfect this aspect, we have developed a marker gene database, PscOA, based on original analysis of single-cell transcriptome literature in the plant science. PscOA comprises 93,319 marker genes from 199 cell types across 17 tissues of 9 plant species.
PscOA offers BLAST search functionality for potential marker genes based on sequence homology and the SCSA tool for marker-based cell type annotation. As a user-friendly platform that integrates plant single-cell transcriptome resources, PscOA serves as a valuable repository for original analysis results related to single-cell transcriptomics studies in the plant science, particularly in the exploration of cell heterogeneity and functional genetics of specific plant cell types.
2. Web interface
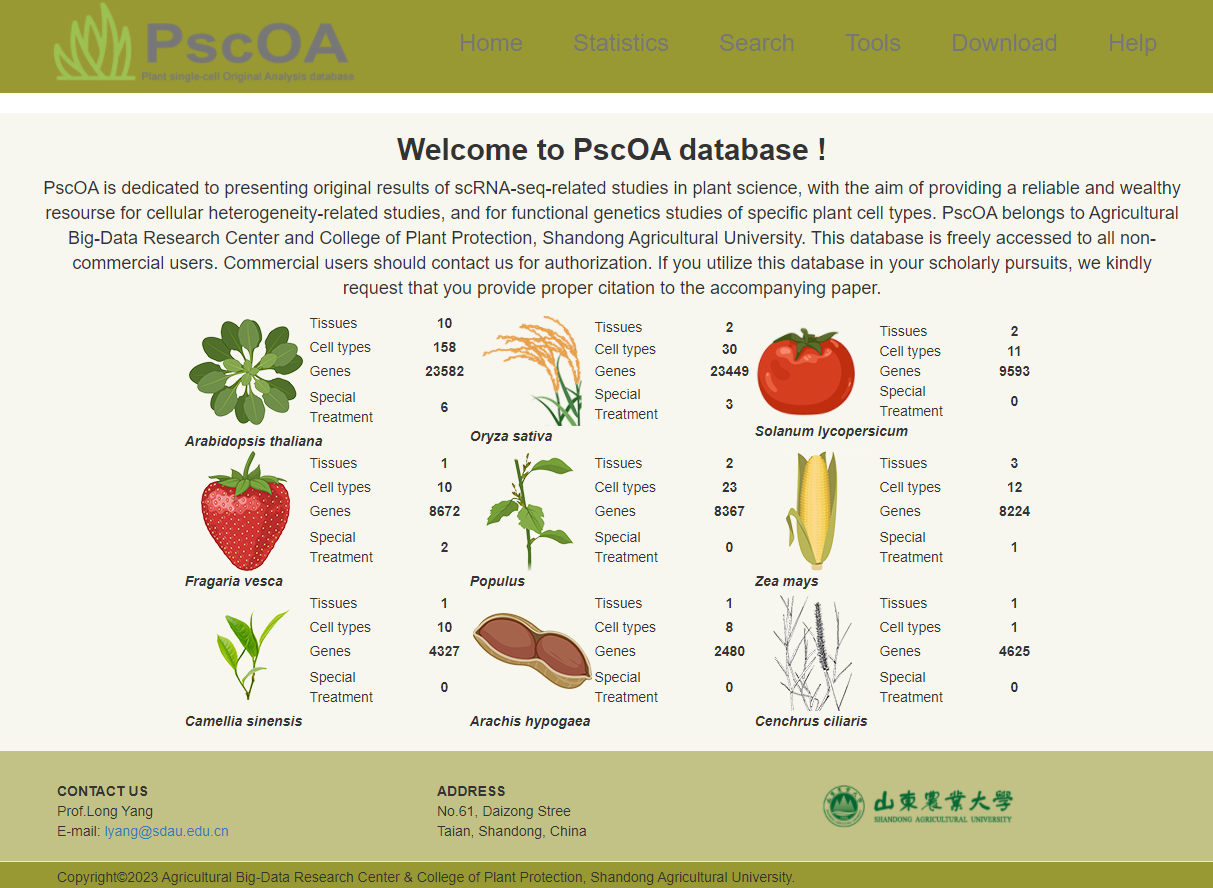
1) Home page
PscOA presents concise statistical information encompassing species, tissues, cell types, genes, as well as special treatment conditions, offering a comprehensive overview for researchers.
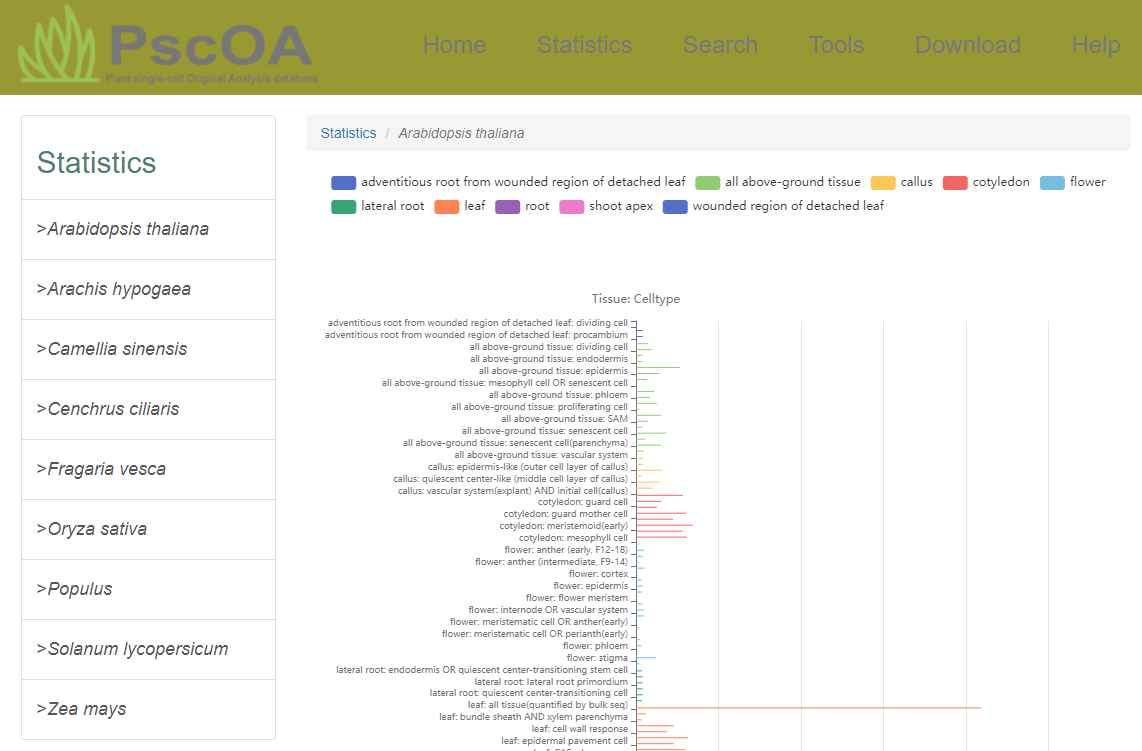
2) Statistics page
PscOA provides detailed statistical information on the number of genes in different tissues and cell types for each species, enabling users to view more comprehensive and in-depth data.
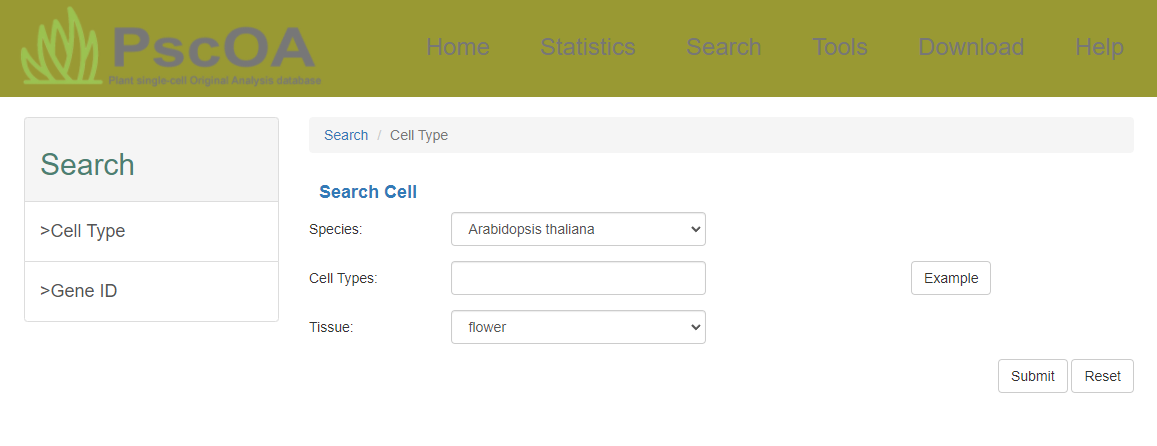
3) Search page
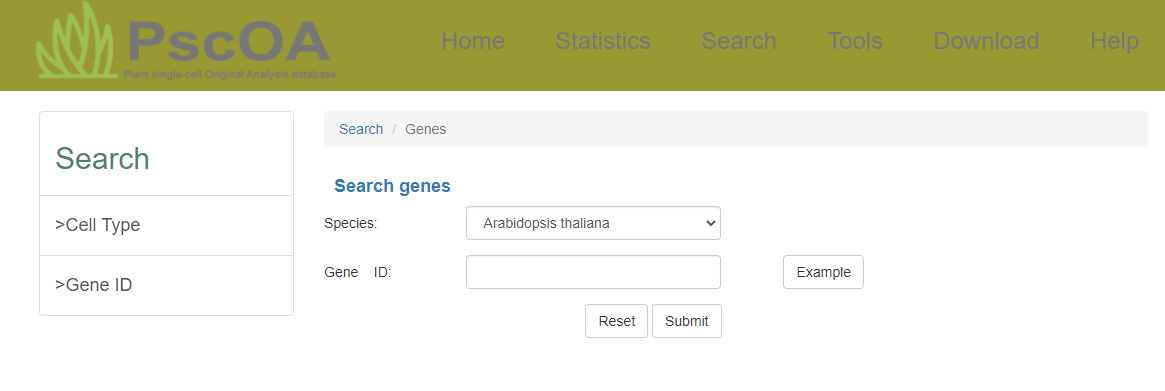
PscOA provides both gene and cell type retrieval options, accompanied by a secondary interface displaying detailed differentially expressed information.
4) Tools page
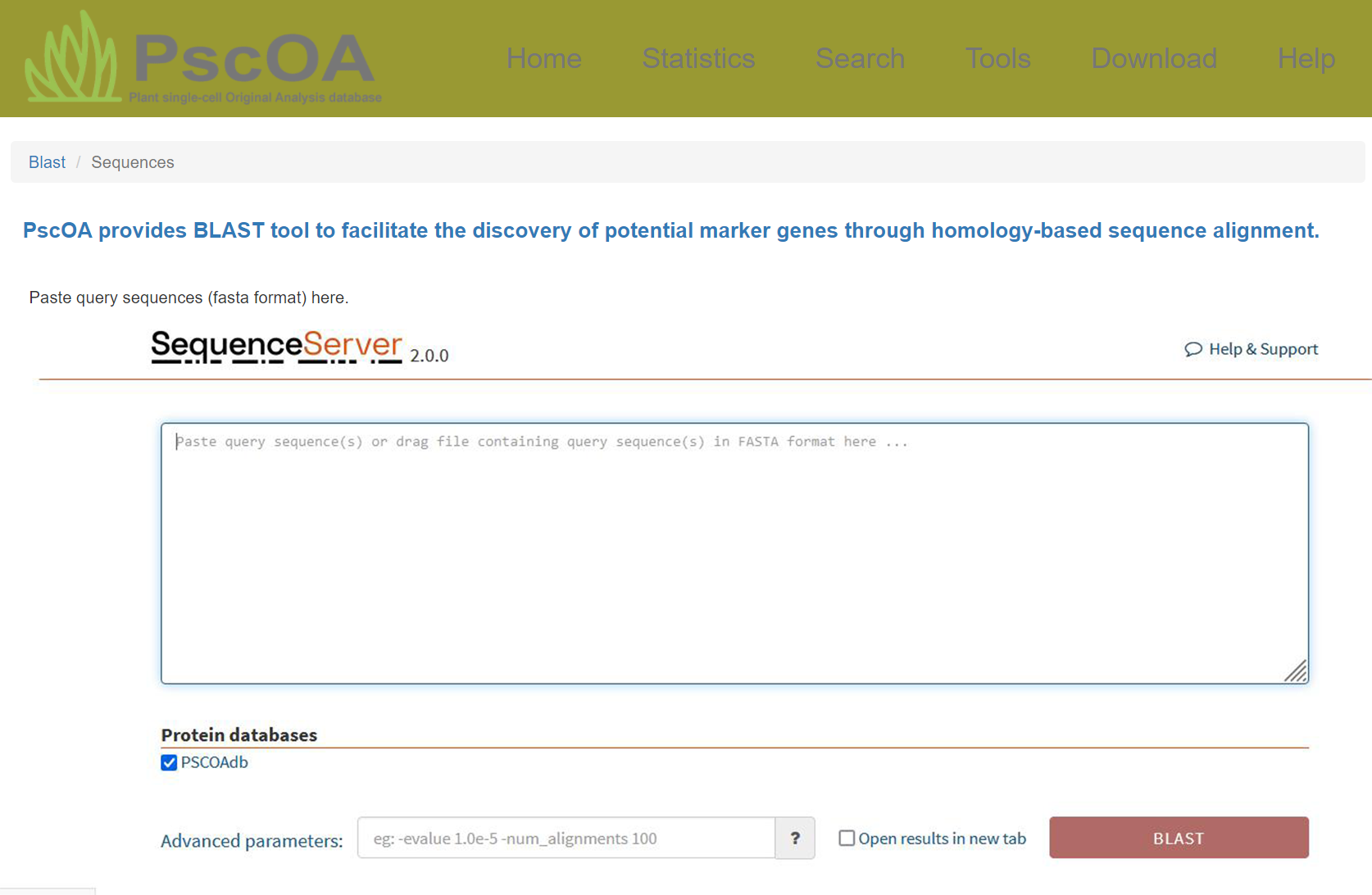
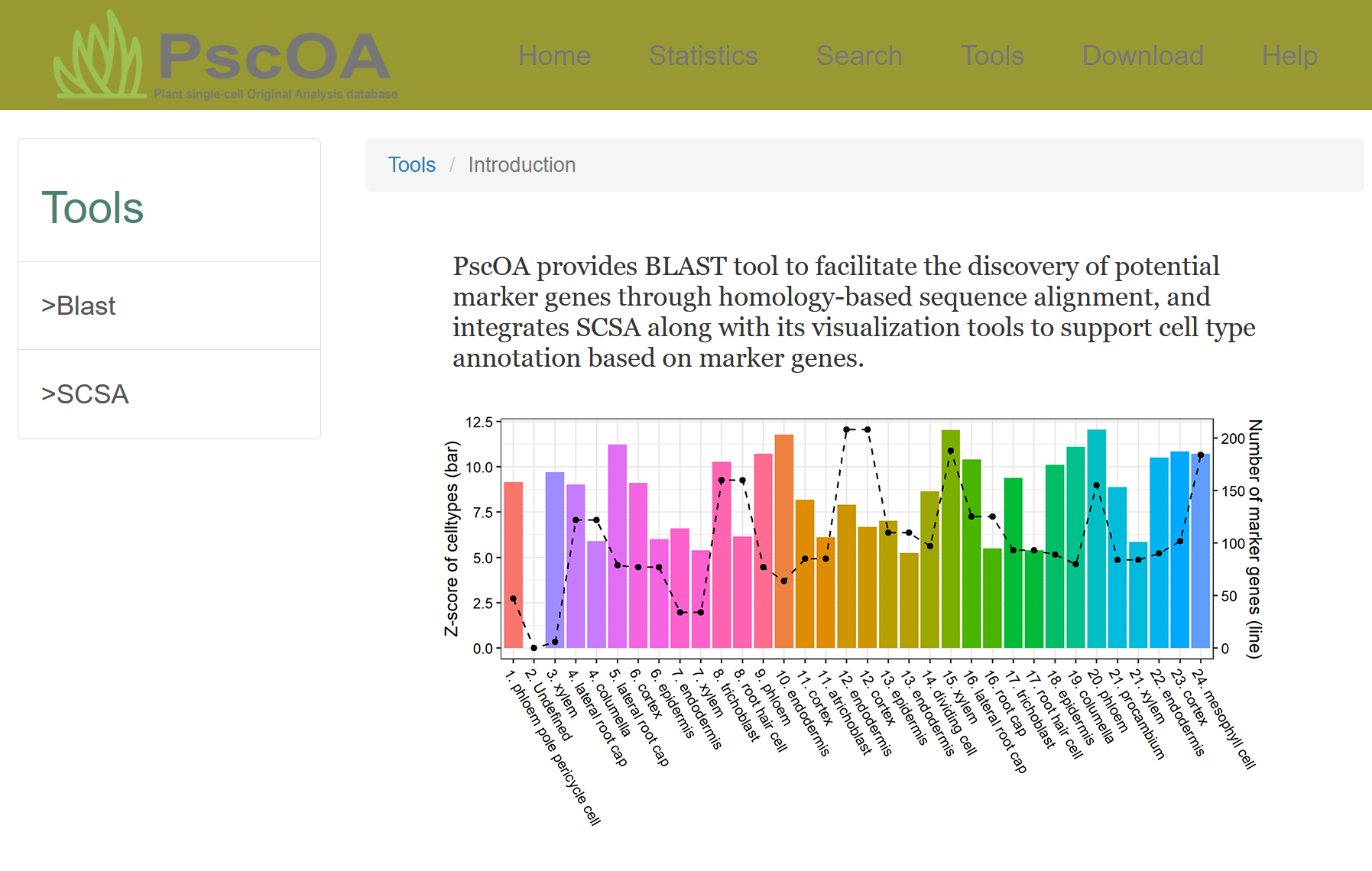
PscOA provides BLAST tool to facilitate the discovery of potential marker genes through homology-based sequence alignment, and integrates SCSA along with its visualization tools to support cell type annotation based on marker genes.
5) Download page
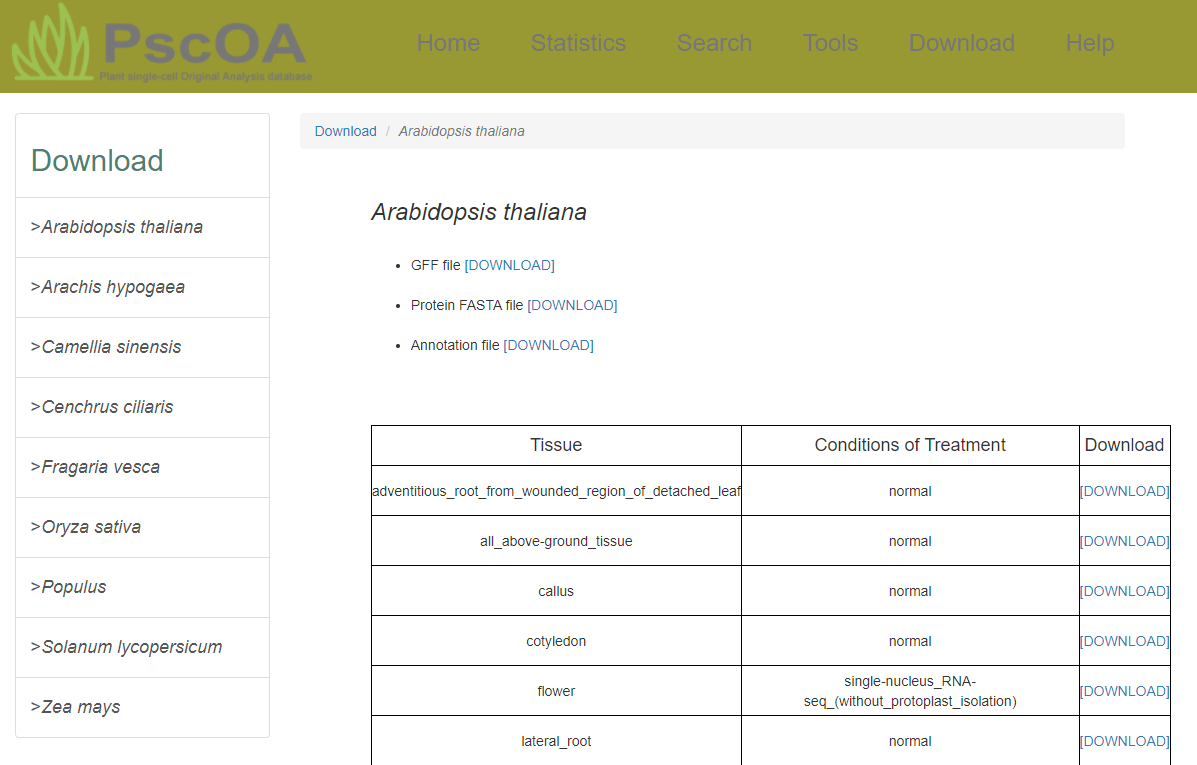
PscOA enables customized downloads based on user-defined parameters to access this database.
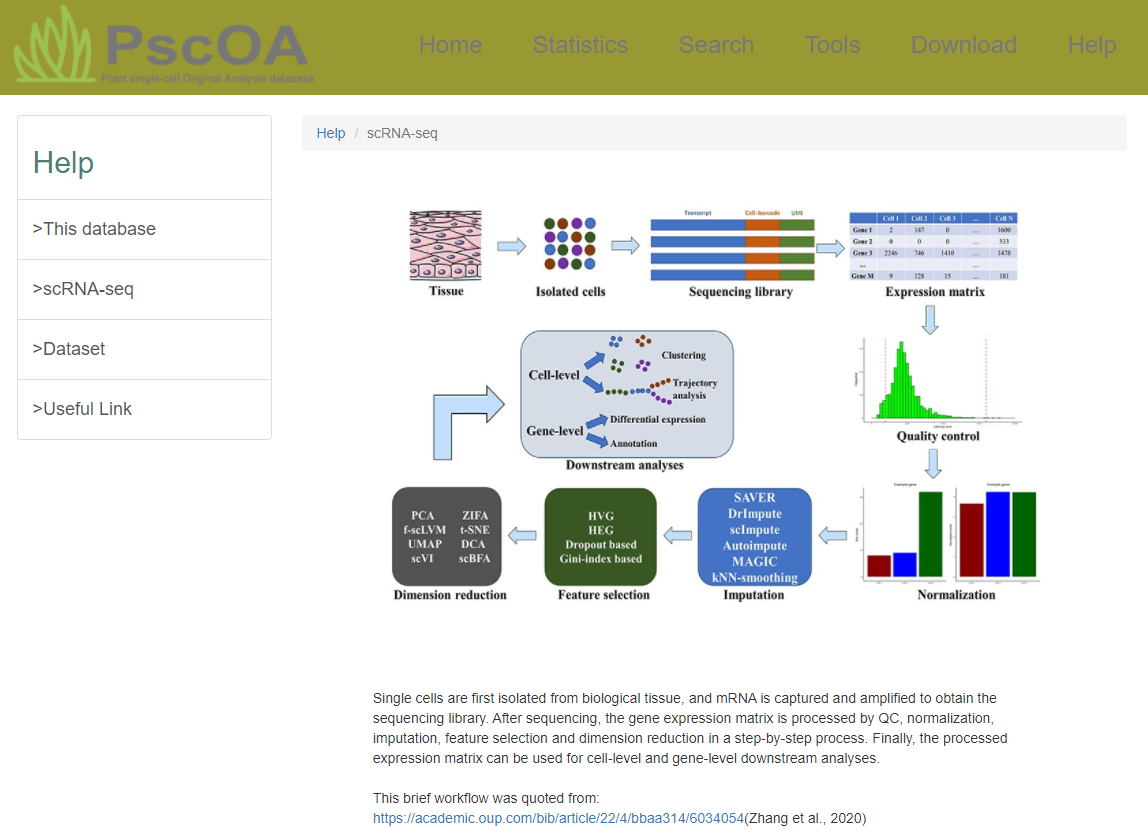
6) Help page
PscOA provides help information about this database, including "This database" subpage to assist users in navigating PscOA smoothly, "scRNA-seq" subpage to take an overview of the scRNA-seq process particularly in plant science, "Dataset" subpage for accessing detailed data sources, and "Useful Link" subpage directing users to bioinformatics-related websites.